HOW WE DO IT
We approach bacteria the same way humans once approached domesticating wild animals. We build an understanding of their needs and abilities, and then we use careful cultivation and genetic engineering to propose a mutually beneficial arrangement.
Wild, Independent Specialists
Bacteria provide nitrogen and phosphorus for the world’s plants. They break down our waste and our livestock’s waste. They produce at least half of the world’s oxygen and can even make clouds in the sky. And they are great at making chemicals.
Bacteria are the undisputed experts in the conversion of biomass to useful chemicals — they taught us how to make many of the chemicals and medicines we now make from petroleum! They are experts in so many processes, and they could teach us so much more, if we could just get to know them better.
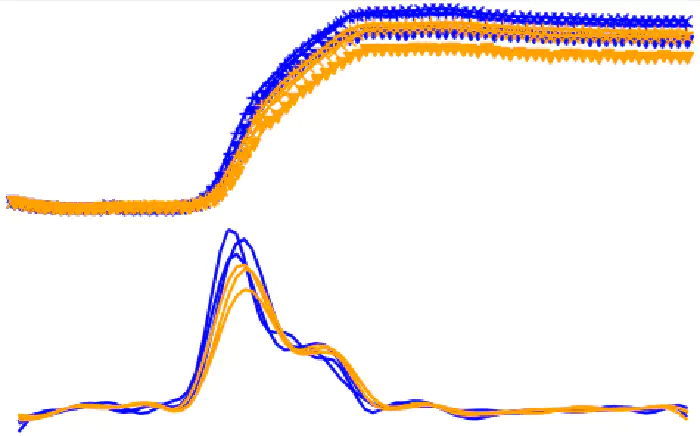
The Right Data Means Less Risk
We cannot domesticate the right partner if we don’t know who has the skills! And the ones with the chops have resisted direct genetic modification to refine their abilities. Our unique platform generates and curates more data about more bacteria more quickly than any other microbiology laboratory. We can tell which species are the most economically relevant to any given industrial process. We can see how to approach them successfully, and which supplementary skills will fully unlock their potential.
As we iterate through generations of genetic engineering, we watch them warm to the task. Our data platform reduces both the risk and time involved in picking the right bacterial partners.

Domestication - A Give and Take
The predominant approach in biotechnology is to transfer genes from skilled bacteria into a genetically manipulable species like E. coli. But asking E. coli to make chemicals or eat plastic is like asking a dog to make milk and eat grass — when a cow is standing right there! Yes, cows and dogs are both four-legged mammals, but a dog needs to eat meat (which is much more expensive than grass) and is going to be miserable (and look weird) hooked up to a milking machine. Making a dog learn how to do a cow’s job just because you’re better at training dogs than cows isn’t leveraging biology. Dogs are the wrong species for the dairy industry, and E. coli is the wrong species for the chemical industry.
We complete an unprecedentedly thorough profile of a bacterium’s needs and wants — and then we use that connection to get it to consent to the minimal genetic engineering needed to make it economically competitive for industrial applications. We don’t fight to move metabolisms into cells that cannot support them at scale. We establish genetic toolkits in cells that already do wonderful chemistries.
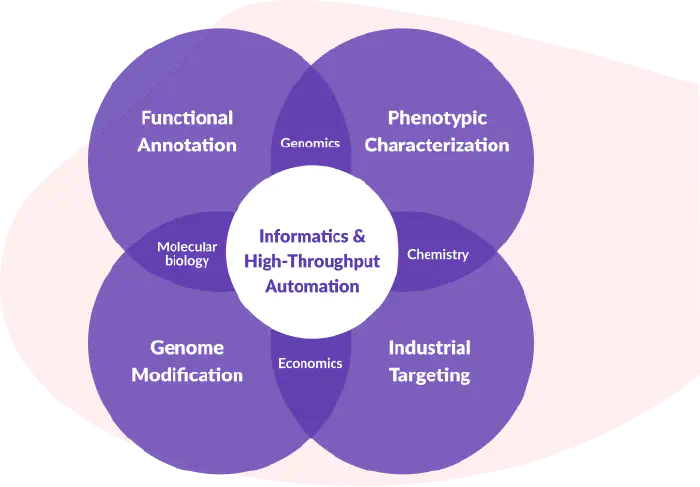
An Ecosystem of Skills
Bacteria are complex systems that navigate even more complex systems. Identifying and exploiting the important details requires skills and techniques from several distinct fields.
We assembled a team of microbiologists, chemists, molecular biologists, data scientists, and computer scientists to automate the collection and curation of bacterial data.